Introduction
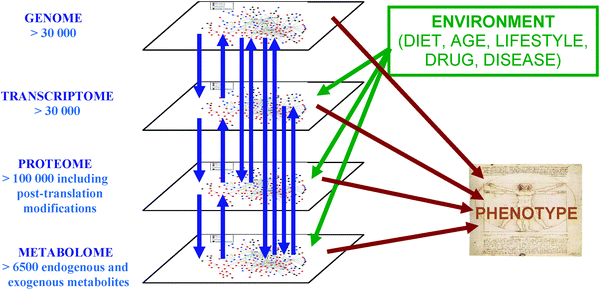
Biological information flows across multiple scales, from individuals to populations, communities, and ecosystems. Metabolomics, the profiling and quantification of metabolites, is a relatively new member of the omics family. Compared with other omics fields, metabolomics focuses on small molecules, typically with molecular weights below 1500 Da. Figure 1.1 illustrates the position of metabolomics within the broader omics landscape[@b.dunn2011].
Metabolomics studies commonly employ GC-MS(Theodoridis et al. 2012; Beale et al. 2018), GC*GC-MS(Tian et al. 2016), LC-MS(Gika et al. 2014), LC-MS/MS(Begou et al. 2017), IM-MS(Levy et al. 2019), infrared ion spectroscopy(Martens et al. 2017), or NMR[@b.dunn2011] to measure metabolites. For broader discussion of analytical methods, see(Zhang et al. 2012), and for an overview of technical progress during 2012-2018, see(Miggiels et al. 2019). This book focuses on mass spectrometry-based metabolomics, especially XC-MS workflows.
Exposomics extends metabolomics toward the comprehensive measurement of environmental, dietary, lifestyle, and xenobiotic exposures together with their biological responses. Because untargeted mass spectrometry is widely used in both fields, exposomics shares many analytical foundations with metabolomics, including sample preparation, feature detection, annotation, normalization, and statistical modeling. However, exposomics also introduces broader questions in exposure science, epidemiology, ethics, and causal interpretation. Therefore, this book focuses on metabolomics workflows, while exposomics is acknowledged as an important related direction beyond the present scope.
Despite its power, metabolomics has several important limitations. No single analytical platform can comprehensively measure the entire metabolome because metabolites differ widely in polarity, abundance, volatility, stability, and ionization behavior. Untargeted studies also face major challenges in confident annotation and identification, while signal intensities can be strongly affected by matrix effects, batch effects, and pre-analytical variation during sample collection, storage, and extraction. Compared with other omics fields, metabolomics usually has lower analytical throughput and higher per-sample experimental complexity, which often limits practical sample size in real studies. As a result, many metabolomics datasets are not large enough to provide stable effect size estimation or strong independent validation, making reproducibility, cross-study comparison, and causal interpretation more difficult.
History
History of Mass Spectrometry
For a broader historical commentary on mass spectrometry, see(Yates Iii 2011). A brief summary is given below:
- 1913, Sir Joseph John Thomson “Rays of Positive Electricity and Their Application to Chemical Analyses.”
Petroleum industry bring mass spectrometry from physics to chemistry
The first commercial mass spectrometer is from Consolidated Engineering Corp to analysis simple gas mixtures from petroleum
In World War II, U.S. use mass spectrometer to separate and enrich isotopes of uranium in Manhattan Project
U.S. also use mass spectrometer for organic compounds during wartime and extend the application of mass spectrometer
1946, TOF, William E. Stephens
1970s, quadrupole mass analyzer
1970s, R. Graham Cooks developed mass-analyzed ion kinetic energy spectrometry, or MIKES to make MRM analysis for multi-stage mass spectrometry
1980s, MALDI rescue TOF and mass spectrometry move into biological application
1990s, Orbitrap mass spectrometry
2010s, Aperture Coding mass spectrometry
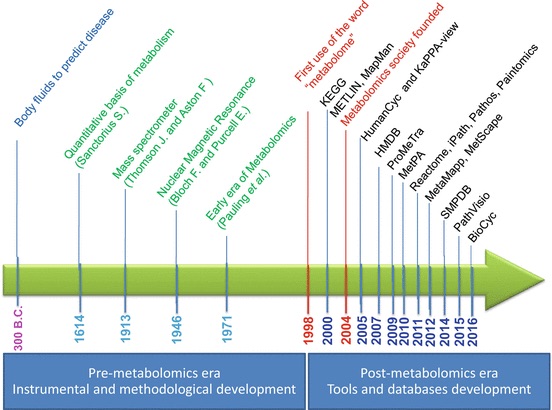
History of Metabolomics
You could check this report(Baker 2011). According to this book section(Kusonmano et al. 2016):
2000-1500 BC some traditional Chinese doctors who began to evaluate the glucose level in urine of diabetic patients using ants
300 BC ancient Egypt and Greece that traditionally determine the urine taste to diagnose human diseases
1913 Joseph John Thomson and Francis William Aston mass spectrometry
1946 Felix Bloch and Edward Purcell Nuclear magnetic resonance
late 1960s chromatographic separation technique
1971 Pauling’s research team “Quantitative Analysis of Urine Vapor and Breath by Gas–Liquid Partition Chromatography”
Willmitzer and his research team pioneer group in metabolomics which suggested the promotion of the metabolomics field and its potential applications from agriculture to medicine and other related areas in the biological sciences
2007 Human Metabolome Project consists of databases of approximately 2500 metabolites, 1200 drugs, and 3500 food components
post-metabolomics era high-throughput analytical techniques
Definition
Metabolomics is actually a comprehensive analysis with identification and quantification of both known and unknown compounds in an unbiased way. Metabolic fingerprinting is working on fast classification of samples based on metabolite data without quantifying or identification of the metabolites. Metabolite profiling always need a pre-defined metabolites list to be quantification(Madsen et al. 2010).
Meanwhile, targeted and untargeted metabolomics are also used in publications. For targeted metabolomics, the majority of the molecules within a biological pathway or a defined group of related metabolites are determined. Sometimes broad collection of known metabolites could also be referred as targeted analysis. Untargeted analysis detect all of possible metabolites unbiased in the samples of interest. A similar concept called non-targeted analysis/screen is actually describe the similar studies or workflow.
Reviews and tutorials
Some nice reviews and tutorials related to this workflow could be found in those papers or directly online:
Glossary
The following abbreviations and terms are used throughout this book:
| ANOVA |
Analysis of Variance |
| APCI |
Atmospheric Pressure Chemical Ionization |
| CID |
Collision-Induced Dissociation |
| DBS |
Dried Blood Spots |
| DDA |
Data-Dependent Acquisition |
| DIA |
Data-Independent Acquisition |
| DoE |
Design of Experiments |
| EIC |
Extracted Ion Chromatogram |
| ESI |
Electrospray Ionization |
| ExWAS |
Exposome-Wide Association Study |
| FDR |
False Discovery Rate |
| GC-MS |
Gas Chromatography-Mass Spectrometry |
| GNPS |
Global Natural Products Social Molecular Networking |
| HILIC |
Hydrophilic Interaction Liquid Chromatography |
| HMDB |
Human Metabolome Database |
| HRMS |
High-Resolution Mass Spectrometry |
| ISF |
In-Source Fragmentation |
| LC-MS |
Liquid Chromatography-Mass Spectrometry |
| ML/DL |
Machine Learning / Deep Learning |
| MR |
Mendelian Randomization |
| MRM |
Multiple Reaction Monitoring |
| MS/MS (MS2) |
Tandem Mass Spectrometry |
| MSI |
Metabolomics Standards Initiative |
| NMR |
Nuclear Magnetic Resonance |
| PCA |
Principal Component Analysis |
| PLS-DA |
Partial Least Squares Discriminant Analysis |
| PMD |
Paired Mass Distance |
| QC |
Quality Control |
| QSPR |
Quantitative Structure-Property Relationship |
| ROI |
Region of Interest |
| RSD |
Relative Standard Deviation |
| SIL-IS |
Stable Isotope-Labeled Internal Standard |
| SPME |
Solid-Phase Microextraction |
| TOF |
Time-of-Flight |
| UPLC |
Ultra-Performance Liquid Chromatography |
| VAMS |
Volumetric Absorptive Microsampling |
Ali, Ahmed, Yasmine Abouleila, Yoshihiro Shimizu, et al. 2019.
“Single-Cell Metabolomics by Mass Spectrometry: Advances, Challenges, and Future Applications.” TrAC Trends in Analytical Chemistry 120 (November): 115436.
https://doi.org/10.1016/j.trac.2019.02.033.
Allam-Ndoul, Bénédicte, Frédéric Guénard, Véronique Garneau, et al. 2016.
“Association Between Metabolite Profiles, Metabolic Syndrome and Obesity Status.” Nutrients 8 (6): 324.
https://doi.org/10.3390/nu8060324.
Alonso, Arnald, Sara Marsal, and Antonio Julià. 2015.
“Analytical Methods in Untargeted Metabolomics: State of the Art in 2015.” Frontiers in Bioengineering and Biotechnology 3 (March).
https://doi.org/10.3389/fbioe.2015.00023.
Baker, Monya. 2011.
“Metabolomics: From Small Molecules to Big Ideas.” Nature Methods 8 (2): 117–21.
https://doi.org/10.1038/nmeth0211-117.
Barnes, Stephen, H. Paul Benton, Krista Casazza, et al. 2016a.
“Training in Metabolomics Research. I. Designing the Experiment, Collecting and Extracting Samples and Generating Metabolomics Data.” Journal of Mass Spectrometry 51 (7): 461–75.
https://doi.org/10.1002/jms.3782.
Barnes, Stephen, H. Paul Benton, Krista Casazza, et al. 2016b.
“Training in Metabolomics Research. II. Processing and Statistical Analysis of Metabolomics Data, Metabolite Identification, Pathway Analysis, Applications of Metabolomics and Its Future.” Journal of Mass Spectrometry 51 (8): 535–48.
https://doi.org/10.1002/jms.3780.
Beale, David J., Farhana R. Pinu, Konstantinos A. Kouremenos, et al. 2018.
“Review of Recent Developments in GC–MS Approaches to Metabolomics-Based Research.” Metabolomics 14 (11): 152.
https://doi.org/10.1007/s11306-018-1449-2.
Begou, O., H. G. Gika, I. D. Wilson, and G. Theodoridis. 2017.
“Hyphenated MS-based Targeted Approaches in Metabolomics.” Analyst 142 (17): 3079–100.
https://doi.org/10.1039/C7AN00812K.
Bilbao, Aivett, Emmanuel Varesio, Jeremy Luban, et al. 2015.
“Processing Strategies and Software Solutions for Data-Independent Acquisition in Mass Spectrometry.” PROTEOMICS 15 (5-6): 964–80.
https://doi.org/10.1002/pmic.201400323.
Broeckling, Corey D., Richard D. Beger, Leo L. Cheng, et al. 2023.
“Current Practices in LC-MS Untargeted Metabolomics: A Scoping Review on the Use of Pooled Quality Control Samples.” Analytical Chemistry 95 (51): 18645–54.
https://doi.org/10.1021/acs.analchem.3c02924.
Bundy, Jacob G., Matthew P. Davey, and Mark R. Viant. 2009.
“Environmental Metabolomics: A Critical Review and Future Perspectives.” Metabolomics 5 (1): 3.
https://doi.org/10.1007/s11306-008-0152-0.
Cajka, Tomas, and Oliver Fiehn. 2016b.
“Toward Merging Untargeted and Targeted Methods in Mass Spectrometry-Based Metabolomics and Lipidomics.” Analytical Chemistry 88 (1): 524–45.
https://doi.org/10.1021/acs.analchem.5b04491.
Castro-Puyana, María, Raquel Pérez-Míguez, Lidia Montero, and Miguel Herrero. 2017.
“Application of Mass Spectrometry-Based Metabolomics Approaches for Food Safety, Quality and Traceability.” TrAC Trends in Analytical Chemistry 93 (August): 102–18.
https://doi.org/10.1016/j.trac.2017.05.004.
Chang, Hui-Yin, Sean M. Colby, Xiuxia Du, et al. 2021.
“A Practical Guide to Metabolomics Software Development.” Analytical Chemistry 93 (4): 1912–23.
https://doi.org/10.1021/acs.analchem.0c03581.
Cheng, Susan, Svati H. Shah, Elizabeth J. Corwin, et al. 2017.
“Potential Impact and Study Considerations of Metabolomics in Cardiovascular Health and Disease: A Scientific Statement From the American Heart Association.” Circulation: Cardiovascular Genetics 10 (2): e000032.
https://doi.org/10.1161/HCG.0000000000000032.
Climaco Pinto, Rui, Ibrahim Karaman, Matthew R. Lewis, et al. 2022.
“Finding Correspondence Between Metabolomic Features in Untargeted Liquid Chromatography–Mass Spectrometry Metabolomics Datasets.” Analytical Chemistry 94 (14): 5493–503.
https://doi.org/10.1021/acs.analchem.1c03592.
Considine, E. C., G. Thomas, A. L. Boulesteix, A. S. Khashan, and L. C. Kenny. 2017.
“Critical Review of Reporting of the Data Analysis Step in Metabolomics.” Metabolomics 14 (1): 7.
https://doi.org/10.1007/s11306-017-1299-3.
Domingo-Almenara, Xavier, J. Rafael Montenegro-Burke, H. Paul Benton, and Gary Siuzdak. 2018.
“Annotation: A Computational Solution for Streamlining Metabolomics Analysis.” Analytical Chemistry 90 (1): 480–89.
https://doi.org/10.1021/acs.analchem.7b03929.
Dryden, Michael D. M., Ryan Fobel, Christian Fobel, and Aaron R. Wheeler. 2017.
“Upon the Shoulders of Giants: Open-Source Hardware and Software in Analytical Chemistry.” Analytical Chemistry 89 (8): 4330–38.
https://doi.org/10.1021/acs.analchem.7b00485.
Du, Xinsong, Juan J. Aristizabal-Henao, Timothy J. Garrett, Mathias Brochhausen, William R. Hogan, and Dominick J. Lemas. 2022.
“A Checklist for Reproducible Computational Analysis in Clinical Metabolomics Research.” Metabolites 12 (1): 87.
https://doi.org/10.3390/metabo12010087.
Dudzik, Danuta, Cecilia Barbas-Bernardos, Antonia García, and Coral Barbas. 2018.
“Quality Assurance Procedures for Mass Spectrometry Untargeted Metabolomics. A Review.” Journal of Pharmaceutical and Biomedical Analysis, Review issue 2017, vol. 147 (January): 149–73.
https://doi.org/10.1016/j.jpba.2017.07.044.
Fenaille, François, Pierre Barbier Saint-Hilaire, Kathleen Rousseau, and Christophe Junot. 2017.
“Data Acquisition Workflows in Liquid Chromatography Coupled to High Resolution Mass Spectrometry-Based Metabolomics: Where Do We Stand?” Journal of Chromatography A 1526 (Supplement C): 1–12.
https://doi.org/10.1016/j.chroma.2017.10.043.
Fessenden, Marissa. 2016.
“Metabolomics: Small Molecules, Single Cells.” Nature 540 (7631): 153–55.
https://doi.org/10.1038/540153a.
Fiehn, Oliver. 2002.
“Metabolomics – the Link Between Genotypes and Phenotypes.” Plant Molecular Biology 48 (1): 155–71.
https://doi.org/10.1023/A:1013713905833.
Gika, Helen G., Georgios A. Theodoridis, Robert S. Plumb, and Ian D. Wilson. 2014.
“Current Practice of Liquid Chromatography–Mass Spectrometry in Metabolomics and Metabonomics.” Journal of Pharmaceutical and Biomedical Analysis, Review
Papers on
Pharmaceutical and
Biomedical Analysis 2013, vol. 87 (January): 12–25.
https://doi.org/10.1016/j.jpba.2013.06.032.
Goldansaz, Seyed Ali, An Chi Guo, Tanvir Sajed, Michael A. Steele, Graham S. Plastow, and David S. Wishart. 2017.
“Livestock Metabolomics and the Livestock Metabolome: A Systematic Review.” PLOS ONE 12 (5): e0177675.
https://doi.org/10.1371/journal.pone.0177675.
González, Oskar, Anne-Charlotte Dubbelman, and Thomas Hankemeier. 2022.
“Postcolumn Infusion as a Quality Control Tool for LC-MS-Based Analysis.” Postcolumn Infusion as a Quality Control Tool for LC-MS-Based Analysis, ahead of print, April.
https://doi.org/10.1021/jasms.2c00022.
González-Domínguez, Álvaro, Núria Estanyol-Torres, Carl Brunius, Rikard Landberg, and Raúl González-Domínguez. 2024.
“QComics: Recommendations and Guidelines for Robust, Easily Implementable and Reportable Quality Control of Metabolomics Data.” Analytical Chemistry 96 (3): 1064–72.
https://doi.org/10.1021/acs.analchem.3c03660.
González-Riano, Carolina, Danuta Dudzik, Antonia Garcia, et al. 2020.
“Recent Developments Along the Analytical Process for Metabolomics Workflows.” Analytical Chemistry 92 (1): 203–26.
https://doi.org/10.1021/acs.analchem.9b04553.
Griffiths, William J., Therese Koal, Yuqin Wang, Matthias Kohl, David P. Enot, and Hans-Peter Deigner. 2010.
“Targeted Metabolomics for Biomarker Discovery.” Angewandte Chemie International Edition 49 (32): 5426–45.
https://doi.org/10.1002/anie.200905579.
Gromski, Piotr S., Howbeer Muhamadali, David I. Ellis, et al. 2015.
“A Tutorial Review: Metabolomics and Partial Least Squares-Discriminant Analysis – a Marriage of Convenience or a Shotgun Wedding.” Analytica Chimica Acta 879 (June): 10–23.
https://doi.org/10.1016/j.aca.2015.02.012.
Hansen, Rebecca L., and Young Jin Lee. 2018.
“High-Spatial Resolution Mass Spectrometry Imaging: Toward Single Cell Metabolomics in Plant Tissues.” The Chemical Record 18 (1): 65–77.
https://doi.org/10.1002/tcr.201700027.
Hites, Ronald A., and Karl J. Jobst. 2018.
“Is Nontargeted Screening Reproducible?” Environmental Science & Technology 52 (21): 11975–76.
https://doi.org/10.1021/acs.est.8b05671.
Jones, Dean P., Youngja Park, and Thomas R. Ziegler. 2012.
“Nutritional Metabolomics: Progress in Addressing Complexity in Diet and Health.” Annual Review of Nutrition 32 (1): 183–202.
https://doi.org/10.1146/annurev-nutr-072610-145159.
Jorge, Tiago F., Ana T. Mata, and Carla António. 2016.
“Mass Spectrometry as a Quantitative Tool in Plant Metabolomics.” Phil. Trans. R. Soc. A 374 (2079): 20150370.
https://doi.org/10.1098/rsta.2015.0370.
Kapoore, Rahul Vijay, and Seetharaman Vaidyanathan. 2016.
“Towards Quantitative Mass Spectrometry-Based Metabolomics in Microbial and Mammalian Systems.” Phil. Trans. R. Soc. A 374 (2079): 20150363.
https://doi.org/10.1098/rsta.2015.0363.
Kennedy, Adam D., Bryan M. Wittmann, Anne M. Evans, et al. 2018.
“Metabolomics in the Clinic: A Review of the Shared and Unique Features of Untargeted Metabolomics for Clinical Research and Clinical Testing.” Journal of Mass Spectrometry 53 (11): 1143–54.
https://doi.org/10.1002/jms.4292.
Kusonmano, Kanthida, Wanwipa Vongsangnak, and Pramote Chumnanpuen. 2016.
“Informatics for Metabolomics.” In
Translational Biomedical Informatics. Advances in
Experimental Medicine and
Biology. Springer, Singapore.
https://doi.org/10.1007/978-981-10-1503-8_5.
Levy, Allison J., Nicholas R. Oranzi, Atiye Ahmadireskety, Robin H. J. Kemperman, Michael S. Wei, and Richard A. Yost. 2019.
“Recent Progress in Metabolomics Using Ion Mobility-Mass Spectrometry.” TrAC Trends in Analytical Chemistry 116 (July): 274–81.
https://doi.org/10.1016/j.trac.2019.05.001.
Liu, Xinyu, Lina Zhou, Xianzhe Shi, and Guowang Xu. 2019.
“New Advances in Analytical Methods for Mass Spectrometry-Based Large-Scale Metabolomics Study.” TrAC Trends in Analytical Chemistry 121 (December): 115665.
https://doi.org/10.1016/j.trac.2019.115665.
Lu, Wenyun, Bryson D. Bennett, and Joshua D. Rabinowitz. 2008.
“Analytical Strategies for LC–MS-based Targeted Metabolomics.” Journal of Chromatography B, Hyphenated
Techniques for
Global Metabolite Profiling, vol. 871 (2): 236–42.
https://doi.org/10.1016/j.jchromb.2008.04.031.
Lu, Xin, and Guowang Xu. 2008.
“LC-MS Metabonomics Methodology in Biomarker Discovery.” In
Biomarker Methods in Drug Discovery and Development, edited by Feng Wang. Methods in
Pharmacology and
Toxicology™. Humana Press.
https://doi.org/10.1007/978-1-59745-463-6_14.
Lv, Wangjie, Zhongda Zeng, Yuqing Zhang, et al. 2022.
“Comprehensive Metabolite Quantitative Assay Based on Alternate Metabolomics and Lipidomics Analyses.” Analytica Chimica Acta 1215 (July): 339979.
https://doi.org/10.1016/j.aca.2022.339979.
Madsen, Rasmus, Torbjörn Lundstedt, and Johan Trygg. 2010.
“Chemometrics in Metabolomics—A Review in Human Disease Diagnosis.” Analytica Chimica Acta 659 (1): 23–33.
https://doi.org/10.1016/j.aca.2009.11.042.
Mangul, Serghei, Thiago Mosqueiro, Richard J. Abdill, et al. 2019.
“Challenges and Recommendations to Improve the Installability and Archival Stability of Omics Computational Tools.” PLOS Biology 17 (6): e3000333.
https://doi.org/10.1371/journal.pbio.3000333.
Martens, Jonathan, Giel Berden, Rianne E. van Outersterp, et al. 2017.
“Molecular Identification in Metabolomics Using Infrared Ion Spectroscopy.” Scientific Reports 7 (June).
https://doi.org/10.1038/s41598-017-03387-4.
Matich, Eryn K., Nita G. Chavez Soria, Diana S. Aga, and G. Ekin Atilla-Gokcumen. 2019.
“Applications of Metabolomics in Assessing Ecological Effects of Emerging Contaminants and Pollutants on Plants.” Journal of Hazardous Materials 373 (July): 527–35.
https://doi.org/10.1016/j.jhazmat.2019.02.084.
Miggiels, Paul, Bert Wouters, Gerard J. P. van Westen, Anne-Charlotte Dubbelman, and Thomas Hankemeier. 2019.
“Novel Technologies for Metabolomics: More for Less.” TrAC Trends in Analytical Chemistry 120 (November): 115323.
https://doi.org/10.1016/j.trac.2018.11.021.
Misra, Biswapriya B. 2018.
“New Tools and Resources in Metabolomics: 2016–2017.” ELECTROPHORESIS 39 (7): 909–23.
https://doi.org/10.1002/elps.201700441.
Misra, Biswapriya B., Johannes F. Fahrmann, and Dmitry Grapov. 2017.
“Review of Emerging Metabolomic Tools and Resources: 2015–2016.” ELECTROPHORESIS 38 (18): 2257–74.
https://doi.org/10.1002/elps.201700110.
Misra, Biswapriya B., and Justin J. J. van der Hooft. 2016.
“Updates in Metabolomics Tools and Resources: 2014–2015.” ELECTROPHORESIS 37 (1): 86–110.
https://doi.org/10.1002/elps.201500417.
Müller, Manfred J., and Anja Bosy-Westphal. 2020.
“From a ‘Metabolomics Fashion’ to a Sound Application of Metabolomics in Research on Human Nutrition.” European Journal of Clinical Nutrition 74 (12): 1619–29.
https://doi.org/10.1038/s41430-020-00781-6.
Ni, Zhixu, Michele Wölk, Geoff Jukes, et al. 2022.
“Guiding the Choice of Informatics Software and Tools for Lipidomics Research Applications.” Nature Methods, December, 1–12.
https://doi.org/10.1038/s41592-022-01710-0.
Pezzatti, Julian, Julien Boccard, Santiago Codesido, et al. 2020.
“Implementation of Liquid Chromatography–High Resolution Mass Spectrometry Methods for Untargeted Metabolomic Analyses of Biological Samples: A Tutorial.” Analytica Chimica Acta 1105 (April): 28–44.
https://doi.org/10.1016/j.aca.2019.12.062.
Place, Benjamin J., Elin M. Ulrich, Jonathan K. Challis, et al. 2021.
“An Introduction to the Benchmarking and Publications for Non-Targeted Analysis Working Group.” Analytical Chemistry 93 (49): 16289–96.
https://doi.org/10.1021/acs.analchem.1c02660.
Rey-Stolle, Fernanda, Danuta Dudzik, Carolina Gonzalez-Riano, et al. 2022.
“Low and High Resolution Gas Chromatography-Mass Spectrometry for Untargeted Metabolomics: A Tutorial.” Analytica Chimica Acta 1210 (June): 339043.
https://doi.org/10.1016/j.aca.2021.339043.
Sarpe, Vladimir, and David C Schriemer. 2017.
“Supporting Metabolomics with Adaptable Software: Design Architectures for the End-User.” Current Opinion in Biotechnology, Analytical biotechnology, vol. 43 (February): 110–17.
https://doi.org/10.1016/j.copbio.2016.11.001.
Schrimpe-Rutledge, Alexandra C., Simona G. Codreanu, Stacy D. Sherrod, and John A. McLean. 2016.
“Untargeted Metabolomics Strategies—Challenges and Emerging Directions.” Journal of The American Society for Mass Spectrometry 27 (12): 1897–905.
https://doi.org/10.1007/s13361-016-1469-y.
Schymanski, Emma L., and Antony J. Williams. 2017.
“Open Science for Identifying ‘Known Unknown’ Chemicals.” Environmental Science & Technology 51 (10): 5357–59.
https://doi.org/10.1021/acs.est.7b01908.
Siskos, Alexandros P., Pooja Jain, Werner Römisch-Margl, et al. 2017.
“Interlaboratory Reproducibility of a Targeted Metabolomics Platform for Analysis of Human Serum and Plasma.” Analytical Chemistry 89 (1): 656–65.
https://doi.org/10.1021/acs.analchem.6b02930.
Smirnov, Kirill S., Tanja V. Maier, Alesia Walker, et al. 2016.
“Challenges of Metabolomics in Human Gut Microbiota Research.” International Journal of Medical Microbiology, Intestinal microbiota - a microbial ecosystem at the edge between immune homeostasis and inflammation, vol. 306 (5): 266–79.
https://doi.org/10.1016/j.ijmm.2016.03.006.
Spicer, Rachel, Reza M. Salek, Pablo Moreno, Daniel Cañueto, and Christoph Steinbeck. 2017.
“Navigating Freely-Available Software Tools for Metabolomics Analysis.” Metabolomics 13 (9).
https://doi.org/10.1007/s11306-017-1242-7.
Spratlin, Jennifer L., Natalie J. Serkova, and S. Gail Eckhardt. 2009.
“Clinical Applications of Metabolomics in Oncology: A Review.” Clinical Cancer Research 15 (2): 431–40.
https://doi.org/10.1158/1078-0432.CCR-08-1059.
Sumner, Lloyd W., Alexander Amberg, Dave Barrett, et al. 2007.
“Proposed Minimum Reporting Standards for Chemical Analysis Chemical Analysis Working Group (CAWG) Metabolomics Standards Initiative (MSI).” Metabolomics : Official Journal of the Metabolomic Society 3 (3): 211–21.
https://doi.org/10.1007/s11306-007-0082-2.
Sumner, Lloyd W, Pedro Mendes, and Richard A Dixon. 2003.
“Plant Metabolomics: Large-Scale Phytochemistry in the Functional Genomics Era.” Phytochemistry, Plant
Metabolomics, vol. 62 (6): 817–36.
https://doi.org/10.1016/S0031-9422(02)00708-2.
Tang, Yanan, Caley B. Craven, Nicholas J. P. Wawryk, Junlang Qiu, Feng Li, and Xing-Fang Li. 2020.
“Advances in Mass Spectrometry-Based Omics Analysis of Trace Organics in Water.” TrAC Trends in Analytical Chemistry 128 (July): 115918.
https://doi.org/10.1016/j.trac.2020.115918.
Theodoridis, Georgios A., Helen G. Gika, Elizabeth J. Want, and Ian D. Wilson. 2012.
“Liquid Chromatography–Mass Spectrometry Based Global Metabolite Profiling: A Review.” Analytica Chimica Acta 711 (January): 7–16.
https://doi.org/10.1016/j.aca.2011.09.042.
Tian, Tze-Feng, San-Yuan Wang, Tien-Chueh Kuo, et al. 2016.
“Web Server for Peak Detection, Baseline Correction, and Alignment in Two-Dimensional Gas Chromatography Mass Spectrometry-Based Metabolomics Data.” Analytical Chemistry 88 (21): 10395–403.
https://doi.org/10.1021/acs.analchem.6b00755.
Uppal, Karan, Douglas I. Walker, Ken Liu, Shuzhao Li, Young-Mi Go, and Dean P. Jones. 2016.
“Computational Metabolomics: A Framework for the Million Metabolome.” Chemical Research in Toxicology 29 (12): 1956–75.
https://doi.org/10.1021/acs.chemrestox.6b00179.
Verhoeven, Aswin, Martin Giera, and Oleg A. Mayboroda. 2020.
“Scientific Workflow Managers in Metabolomics: An Overview.” Analyst 145 (11): 3801–8.
https://doi.org/10.1039/D0AN00272K.
Viant, Mark R., Timothy M. D. Ebbels, Richard D. Beger, et al. 2019.
“Use Cases, Best Practice and Reporting Standards for Metabolomics in Regulatory Toxicology.” Nature Communications 10 (1): 3041.
https://doi.org/10.1038/s41467-019-10900-y.
Vinaixa, Maria, Emma L. Schymanski, Steffen Neumann, Miriam Navarro, Reza M. Salek, and Oscar Yanes. 2016.
“Mass Spectral Databases for LC/MS- and GC/MS-based Metabolomics: State of the Field and Future Prospects.” TrAC Trends in Analytical Chemistry 78 (April): 23–35.
https://doi.org/10.1016/j.trac.2015.09.005.
Vitale, Chiara Maria, Arjen Lommen, Carolin Huber, et al. 2022.
“Harmonized Quality Assurance/Quality Control Provisions for Nontargeted Measurement of Urinary Pesticide Biomarkers in the HBM4EU Multisite SPECIMEn Study.” Analytical Chemistry 94 (22): 7833–43.
https://doi.org/10.1021/acs.analchem.2c00061.
Wallach, Joshua D., Kevin W. Boyack, and John P. A. Ioannidis. 2018.
“Reproducible Research Practices, Transparency, and Open Access Data in the Biomedical Literature, 2015–2017.” PLOS Biology 16 (11): e2006930.
https://doi.org/10.1371/journal.pbio.2006930.
Warth, Benedikt, Scott Spangler, Mingliang Fang, et al. 2017.
“Exposome-Scale Investigations Guided by Global Metabolomics, Pathway Analysis, and Cognitive Computing.” Analytical Chemistry 89 (21): 11505–13.
https://doi.org/10.1021/acs.analchem.7b02759.
Weljie, Aalim M., Jack Newton, Pascal Mercier, Erin Carlson, and Carolyn M. Slupsky. 2006.
“Targeted Profiling: Quantitative Analysis of 1H NMR Metabolomics Data.” Analytical Chemistry 78 (13): 4430–42.
https://doi.org/10.1021/ac060209g.
Wishart, David S. 2016.
“Emerging Applications of Metabolomics in Drug Discovery and Precision Medicine.” Nature Reviews Drug Discovery 15 (7): 473–84.
https://doi.org/10.1038/nrd.2016.32.
Wolfender, Jean-Luc, Guillaume Marti, Aurélien Thomas, and Samuel Bertrand. 2015.
“Current Approaches and Challenges for the Metabolite Profiling of Complex Natural Extracts.” Journal of Chromatography A, Editors’
Choice IX, vol. 1382 (February): 136–64.
https://doi.org/10.1016/j.chroma.2014.10.091.
Yates Iii, John R. 2011.
“A Century of Mass Spectrometry: From Atoms to Proteomes.” Nature Methods 8 (8): 633–37.
https://doi.org/10.1038/nmeth.1659.
Yuan, Min, Susanne B. Breitkopf, Xuemei Yang, and John M. Asara. 2012.
“A Positive/Negative Ion–Switching, Targeted Mass Spectrometry–Based Metabolomics Platform for Bodily Fluids, Cells, and Fresh and Fixed Tissue.” Nature Protocols 7 (5): 872–81.
https://doi.org/10.1038/nprot.2012.024.
Zenobi, R. 2013.
“Single-Cell Metabolomics: Analytical and Biological Perspectives.” Science 342 (6163): 1243259.
https://doi.org/10.1126/science.1243259.
Zhang, Aihua, Hui Sun, Ping Wang, Ying Han, and Xijun Wang. 2012.
“Modern Analytical Techniques in Metabolomics Analysis.” The Analyst 137 (2): 293–300.
https://doi.org/10.1039/C1AN15605E.
Zhou, Juntuo, and Yuxin Yin. 2016.
“Strategies for Large-Scale Targeted Metabolomics Quantification by Liquid Chromatography-Mass Spectrometry.” Analyst 141 (23): 6362–73.
https://doi.org/10.1039/C6AN01753C.